by Buck Institute
August 15, 2019 . Press Release
Uric acid pathologies shorten lifespan in flies, highlighting the need for screening in humans
Backed by human genetics, research in flies provides potential drug targets for gout, metabolic syndrome, diabetes and kidney stones
Few people get their level of uric acid, a breakdown product of metabolism, measured in their blood. Based on Buck research published August 15 in PLOS Genetics, it might be time to rethink that, given that 20 percent of the population have elevated levels of uric acid, increasing their risk for gout, kidney stones, metabolic syndrome, obesity, diabetes and early death.
“People can be at genetic risk for high uric acid and not know it,” says Pankaj Kapahi, PhD, Buck professor and senior author of the paper. “Medical practitioners haven’t been paying sufficient attention to uric acid and perhaps they should.” Kapahi thinks uric acid should be included in routine check-ups similar to those done for cholesterol and blood glucose. “Uric acid levels often go up with age and it’s important for longevity,” he explains. “Gout is also associated with premature mortality in humans. Conversely, studies from the Institute for Aging Research led by Nir Barzilai at the Albert Einstein College of Medicine show that many centenarians are genetically predisposed to having low levels of uric acid.”
While humans lost the gene for metabolizing uric acid about 15 million years ago, most species, including insects, kept it. In this study, researchers in the Kapahi lab “humanized” fruit flies by knocking down the uric acid oxidase gene. The altered flies built up uric acid in their bodies only when triggered by a diet rich in purines, but showed no deleterious effect under dietary restriction. In humans, a high-purine diet is linked to excessive intake of alcohol and red meat. Similarly, sugary beverages are driving factors for the built up of uric acid.
Scientists discovered that the insulin-like signaling (ILS) pathway, which animals use to sense nutrients, plays a role in regulating uric acid levels. Suppressing the ILS pathway lowered uric acid in the genetically altered animals, providing potential targets for new drug development. Additionally, the conserved role of the ILS pathway in modulating uric acid levels was supported by a human gene study which identified variations in two ILS genes associated with serum uric acid levels or gout, the most common inflammatory arthritis.
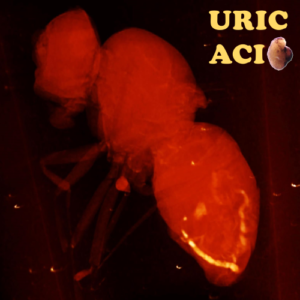
This image shows a micro-CT scan of Drosophila melanogaster with diminished expression of urate oxidase reducing the enzymatic degradation of the aggregation-prone purine intermediate uric acid. Radiolucent structures are shown in red. Radiopaque objects are shown in yellow and demonstrate the formation of uric acid concretions, i.e. events of ectopic biomineralization resembling human uric acid kidney stones. The image replacing the letter “D” in the word uric acid represents a concretion dissected from the same fly.
Sven Lang, a former postdoctoral fellow in the Kapahi lab who led the research, found that a high-purine diet shortened the lifespan of the humanized flies by 48 percent, while suppressing the ILS pathway prevented the rise in uric acid levels even when the flies ate the high yeast diet. The team also found that an increase of free radicals generated by an enzyme called NADPH oxidase (NOX) mediated kidney stone formation and early mortality in the flies. “We were able to inhibit the increase in free radicals using the common antioxidant vitamin C which reduced the burden of kidney stones and improved survival in the animals,” said Lang, who now has his own laboratory at the Department of Medical Biochemistry and Molecular Biology at Saarland University in Homburg, Germany.
“Changes in diet are not always sufficient to bring down levels of uric acid, so it’s important to track it and make sure that patients who need preventive drug treatment get it,” said study co-author Marshall Stoller, MD, head of the urinary stone division at the University of California, San Francisco’s Department of Urology. “In this research we used fruit flies to recapitulate what happens in humans and we discovered new drug targets. It’s our hope that we’ll be able to utilize these targets to develop new drugs for the many diseases linked to uric acid accumulation.”
Citation: A conserved role of the insulin-like signaling pathway in diet-dependent uric acid pathologies in Drosophila melanogaster
DOI: 10.1371/journal.pgen.1008318.
Other Buck researchers involved in the study include Tyler A. Hilsabeck, Kenneth A. Wilson, Amit Sharma, Neelanjan Bose, Jennifer N. Beck, Mark A. Watson, and Arnold Kahn. Other collaborators include Deanna J. Brackman and Kathleen Giacomini, Department of Bioengineering and Therapeutic Sciences, University of California, San Francisco; Ling Chen and Sunita Ho, Division of Biomaterials, University of California, San Francisco; David W. Killilea, Nutrition and Metabolism Center, Children’s Hospital Oakland Research Institute, Oakland, CA; and Thomas Chi, Department of Urology, University of California, San Francisco.
This work was funded by grants from the American Federation of Aging Research and Hillblom foundations and NIH (R01AG038688 & RO1AG045835) granted to Pankaj Kapahi. Development of the RPGEH was supported by The Robert Wood Johnson Foundation, the Wayne and Gladys Valley Foundation, The Ellison Medical Foundation, KPNC, and the Kaiser Permanente National and Regional Community Benefit Programs. Work and personnel for this project were funded by NIH grant R01DK103729. Deanna Brackman was additionally funded by NIH Training Grant T32 GM717537.
Science is showing that while chronological aging is inevitable, biological aging is malleable. There's a part of it that you can fight, and we are getting closer and closer to winning that fight.
Eric Verdin, MD, Buck Institute President and CEO