by Buck Institute
October 21, 2020 . BLOG
Driving that Gene, Mutations are Seen: Spreading a new trait through a viral population
 By Marius Walter, PhD
By Marius Walter, PhD
Postdoctoral Research Fellow in the Verdin lab
The Buck Institute for Research on Aging is best known for its work on, well, aging, but some labs also focus on other related areas. The Verdin lab, where I am a postdoctoral scientist, studies the immune system and aging, but also has scientists focused on virology, which is my field. While infectious diseases are not usually considered age-related, we know for certain that many viruses represent an increased risk for older individuals, as COVID-19 has unfortunately made clear.
In 2016, I was a recent PhD graduate in Paris and very excited about all the novel perspectives enabled by the CRISPR revolution. CRISPR, a naturally occurring defense tool for bacteria, had recently been harnessed as a remarkably simple and efficient way to edit DNA. In fact, this discovery was so remarkable that it just won the 2020 Nobel Prize in Chemistry! In particular, I found the recent development of gene drives fascinating, as they offered the possibility to alter populations of insects in the wild and could help to eradicate malaria and other mosquito-borne diseases.
Gene drives are designed to efficiently spread a genetic modification through a population. They usually rely on CRISPR editing, where an engineered DNA sequence containing the CRISPR system is inserted into the genome of animals. The modified DNA region also harbors a sequence that guides the CRISPR tools to the very same location in the unmodified genome. Sexual reproduction leads to the duplication of the new DNA sequence, ensuring its spread through the population. Because the strategy relies on characteristics unique to sexual reproduction, it had generally been assumed that a gene drive could only be engineered in sexually reproducing organisms such as animals, plants or fungi, excluding bacteria and viruses.
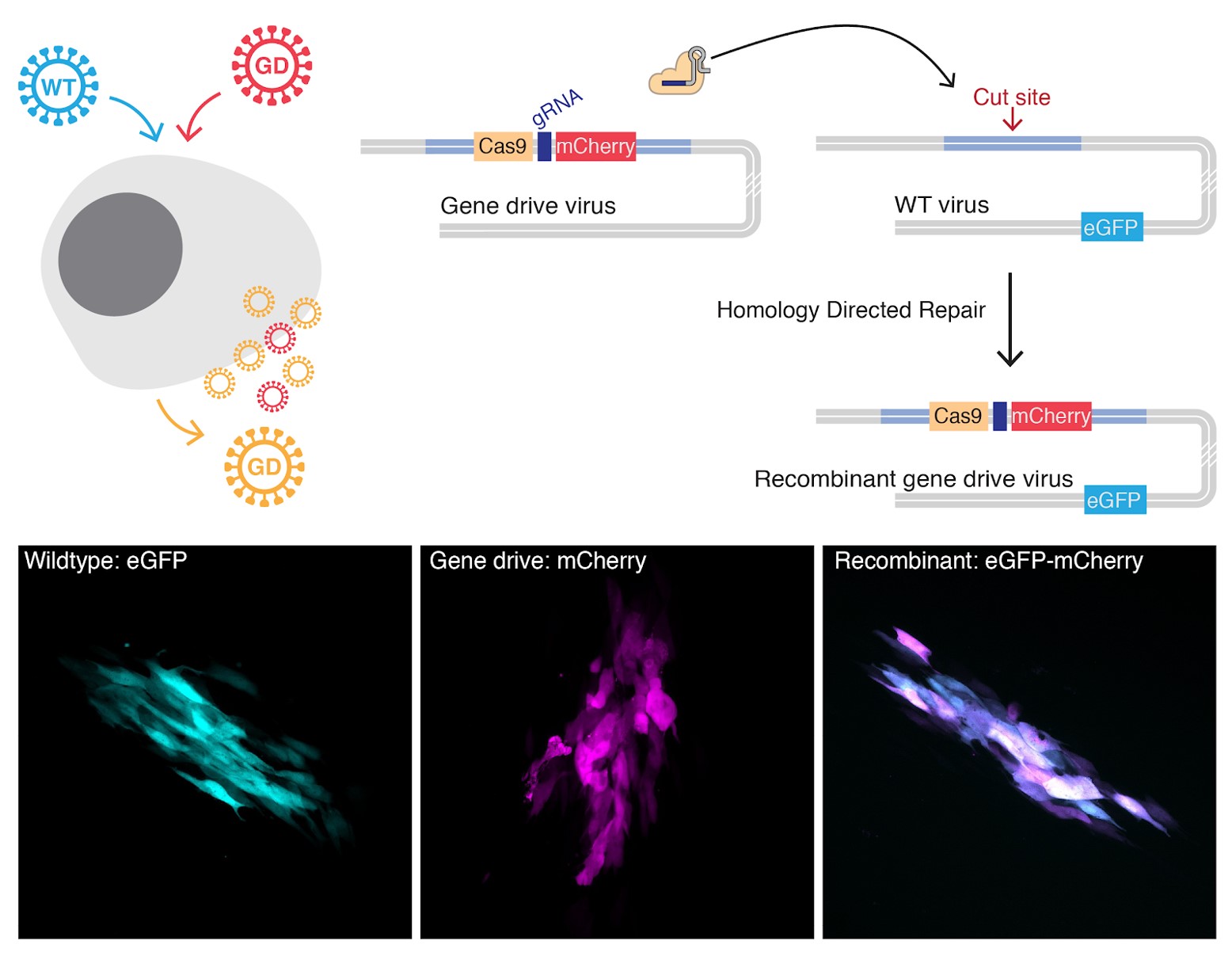
However, as I was preparing for my postdoc interview with the lab of Eric Verdin, I realized that during a viral infection, viruses can accumulate hundreds or thousands of genome copies in infected cells. Additionally, several lines of evidence indicate that cells are frequently co-infected by multiple virions and that the recombination of viral genomes (the exchange of DNA sequences between viruses, by a process similar to what happens during sexual reproduction) is a well-known and widespread source of diversity for many viruses. In particular, herpesviruses are DNA viruses with large double-stranded DNA genomes and frequently undergo recombination during their replication cycle. Upon joining Eric’s lab a few months later, we reasoned that these properties enabled the design of a gene drive strategy that doesn’t involve sexual reproduction, but instead relies on co-infection of a given cell by both a naturally occurring and an engineered virus. Upon co-infection, the unmodified viral genome is cut and the engineered DNA sequence is copied into the natural genome. This results in new gene drive viruses with engineered DNA that can progressively replace the naturally occurring viral population.
In our recent study published in Nature Communications, we presented a proof of principle for such a viral gene drive, using human cytomegalovirus (human herpesvirus 5) as a model. We designed a gene drive that we thought would reduce the infectivity of the virus by editing the DNA to make it less able to reproduce and spread. We demonstrated that gene drive viruses can replace their naturally occurring counterparts and spread in the viral population in cell culture experiments. Moreover, we showed that the infectivity of the modified virus could be drastically reduced, which opens novel therapeutic strategies against herpesviruses.
Herpesviruses are universal pathogens. They establish life-long persistent infections and are implicated directly or indirectly in numerous human diseases: Herpes simplex 1 and 2 for example give painful oral and genital blisters; hCMV is an important threat to immunocompromised patients, such as HIV-infected individuals, receivers of organ transplants and unborn infants; and Epstein-Barr virus or Kaposi’s sarcoma-associated viruses are major causes of cancer. Moreover, because herpesviruses are life-long pathogens, they represent an important risk factor for many age-related diseases. The painful rash that characterizes Shingles in older patients is for example a resurgence of the dormant virus that gives chickenpox in children. Life-long hCMV infection in the elderly has also been associated with chronic inflammation and immunosenescence — the declining function of the immune system with age. Similarly, Roseola, Herpes Simplex and Epstein-Barr viruses have been implicated in Alzheimer’s and other neurodegenerative diseases.Treatment options exist, but drug resistance and adverse secondary effects render the development of new therapeutic solutions necessary. We showed in vitro (in a test tube) that a gene drive virus presenting severe replicative defects could spread into the viral population and ultimately reduce viral levels. Our hope is to ultimately design similar gene drive systems that could be used as treatment against these diseases. Patients infected with an herpesvirus and unable to control it could be superinfected with a gene drive virus that would help to reduce the infection.
While much research is needed before the technology can be adapted to function in an animal model or in humans, we believe that our work bridges the gene drive and virology fields, and represents a new area of research with exciting perspectives. With virology so much in the news right now, it is particularly exciting to be sharing optimistic news about a novel approach to treating dangerous viruses.
- Burt, A. Site-specific selfish genes as tools for the control and genetic engineering of natural populations. Proceedings. Biol. Sci 270, (2003).
- Gantz, V. M. et al. Highly efficient Cas9-mediated gene drive for population modification of the malaria vector mosquito Anopheles stephensi. Proceedings of the National Academy of Sciences 112, E6736–E6743 (2015).
- Walter, M. & Verdin, E. Viral gene drive in herpesviruses. Nat. Commun. 11, 4884 (2020).
SHARE